Every living organism is built from a design …
which is coded in genes. All cells in the human body, except red blood cells, contain a nucleus which holds the genetic information necessary to make a completely new human being. In Sleeper, Woody Allen and Diane Keaton are supposed to clone a dead person from his nose, but the fact is that once the details of human cloning get worked out, only a single cell is going to be necessary to clone a person.
You might consider it pretty dang wasteful that every cell carries the entire genome—all the DNA needed to make a new person. If you’re a white blood cell and your job is to fight infections, why would you carry around the DNA to make ear hairs or produce testosterone? No one knows.
It is helpful to remember that in genetics, things often come in pairs. The DNA in our bodies sits on 46 total chromosomes which are grouped into 23 distinct pairs, numbered 1–23. Your mom supplies one half-set of chromosomes and your dad supplies the other. These are those fuzzy black images that look like Xs and Ys from our high school biology textbooks.
On each chromosome are what we call genes. What is a gene? Well, it is a sequence of DNA that tells the body how to make a particular protein. For example, the insulin gene is located on chromosome 11. Proteins are everything to an organism; eyes, brain, bones, muscle, antibodies, cells, earwax, everything in the body—directly or indirectly—comes from proteins. Bones have calcium, sure, but they have a protein frame which allows the calcium to deposit there. Cell membranes are made of fat, but this fat is manufactured and put in place by proteins. You eat a cheeseburger to get energy, but the chemical reactions which turn a cheeseburger into energy are orchestrated by proteins.
D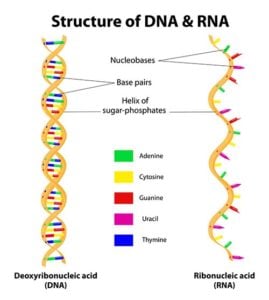 NA mostly consists of a long string of only four different molecules: Adenine, Guanine, Cytosine, and Thymine. They are abbreviated A, G, C, and T. Proteins, such as the protein which makes tendons or nerves, are not made of these molecules. Instead, these molecules are instructions that tell the body how to make a protein. They are best thought of as letters in a written recipe. You don’t eat the word C-H-O-C-O-L-A-T-E in a recipe which calls for chocolate; you don’t eat the ink and paper it is printed on. Instead, those letters tell you what ingredient to grab from the pantry. In the case of DNA, it is exactly the same. A sequence of those four letters tells the body what ingredient to put in each protein.
NA mostly consists of a long string of only four different molecules: Adenine, Guanine, Cytosine, and Thymine. They are abbreviated A, G, C, and T. Proteins, such as the protein which makes tendons or nerves, are not made of these molecules. Instead, these molecules are instructions that tell the body how to make a protein. They are best thought of as letters in a written recipe. You don’t eat the word C-H-O-C-O-L-A-T-E in a recipe which calls for chocolate; you don’t eat the ink and paper it is printed on. Instead, those letters tell you what ingredient to grab from the pantry. In the case of DNA, it is exactly the same. A sequence of those four letters tells the body what ingredient to put in each protein.
The ingredients that make proteins are called amino acids. There are 20 amino acids in the human body, and from these you can assemble any protein you want. Imagine a house: it is made of lumber, drywall, carpet, roofing materials, etc. Amino acids are like the materials which make a house. DNA is the blueprint that indicates what materials, and in what order, should be strung together for construction. Since there are 20 different amino acids, but only four letters in DNA (A,G,C, and T), the letters are read in groups of three to determine which amino acid to choose when building a protein. In English, the letter “B” means nothing, but B-O-Y means something. In living organisms, the sequence GAG indicates that the amino acid glutamic acid should be added to the developing protein. If we know the sequence of letter codes in a gene, we can then easily figure out what is the sequence of amino acids for any protein.
As noted, DNA is located in the nucleus of cells. But what do you call the rest of the cell outside the nucleus? You call it the cytoplasm and that is where proteins are manufactured. In order to get genes into the cytoplasm so that a protein can be manufactured, a special kind of molecule called messenger RNA, which is very similar to DNA, is created and serves as a copy of the gene that floats into the cytoplasm. It is similar to Xeroxing a recipe from a cookbook and working off the copied page so as to not spill butter and milk on the original cookbook.
Imagine you took a human cell and sequenced the entire genetic code. You would have about three billion letters, or base pairs, as they are called. Why the stupid term base pairs when letters is much easier to understand? Well, the reason for that term has to do with how DNA is copied. Every strand of DNA is bound to a complimentary twin strand, like a nut and a bolt, Watson and Crick, or better yet, two pieces of Velcro. Each strand of DNA sits in the nucleus stuck to a partner strand. When the two stands are stuck together, A (Adenine) always is bound to T (Thymine) and C (Cytosine) is always bound to G (Guanine). This creates the so-called “double helix.” The double helix of DNA is just like a spiral staircase where half of each step is one letter, and the other half of the step is its companion. Another visual is to imagine a long strip of Velcro with the two sides stuck together. Just twist the ends a few times and that is what DNA looks like. The advantage of this arrangement is that if you split the staircase down the middle, it is easy to rebuild it into two identical staircases. Here’s an example with English. Suppose you wrote a book and on every line of text, you put the opposite word on the next line. So if the intended line from the book reads, “Children love smiles,” the next line would read, “Adults hate frowns.” Splitting the two in half would allow you to make two whole copies of the originals. Give me “frowns” and I can figure out that it should be paired with “smiles.” Clever, no? This is also how messenger RNA is made. The double helix is unwound and split, and messenger RNA binds to the nonsensical, or “Adults hate frowns” side of DNA, which is called the 3’ (three prime) strand among the cognoscenti. The result is a copy of the original strand of DNA which can float into the cytoplasm and be used as a template to manufacture a protein. What I’ve presented here is just a tiny speck of the DNA story, but it is enough to consider what it means to obtain the DNA sequence of every gene in the body.
There are not that many genes in a person, only 32,000. But imagine that you could take a person and get his or her entire genetic code, that is, list the exact sequence of all the “letters” which make up a particular human being. It would be about three billion letters long, or as long as one thousand copies of War and Peace. However, a typical protein in our body is only 360 amino acids, so there seems to be 10 to 20 times as much DNA in our cells as we need to make all of our proteins.
“Woah,” as Neo might say. What’s going
But this fact is going to save us money and move medicine forward. Because most of the three billion letters in our genome do nothing and can be ignored, if we want to sequence the entire DNA of a person, we can skip 90% of it and just sequence the exons, just the parts of DNA that actually make proteins. This advance is called whole exome sequencing.
For a few thousand dollars, you can have all your exons sequenced. I’m sure the price is going to come down in the next few years. One day in the not too distant future, doctors will have your entire exome on a computer and be able to make clinical decisions based on your genes. A recent article in the New England Journal of Medicine from a research group at Baylor University in Texas rounded up 250 people and performed this process—whole exome sequencing. That is, they took the DNA, but instead of sequencing everything, they ignored the junk which is 90% of DNA and just sequenced the active genes. It was about $7,000 per person. Most of these patients were children with neurological problems, mental retardation, or other birth defects. In 25% of the cases, they made the diagnosis by sequencing the genome. Many of these diagnoses were for extremely rare genetic diseases that most of us, including physicians, have never heard of, such as Floating-Harbor syndrome, which causes facial abnormalities and language delay and has only 50 cases reported in the world’s literature, or KBG syndrome, which causes mental retardation, bone and structural changes, and also causes patients to become international spies.
Genomic medicine means sequencing the DNA of a patient and using this information to guide clinical decision making. Imagine a patient who comes to see me for migraines. Often, it takes trial and error to determine which of the dozen drugs we can use will be best for a particular person. Given a close family member with a similar diagnosis and a good response to a particular medication, it makes sense to try that same medication first on the patient. Some patients don’t respond to typical headache medications such as Imitrex and end up doing extremely well on drugs used to treat seizures or high blood pressure. It seems very likely that if I had all of my patient’s genome sequenced and I could compare it to a database of a million patients who had been treated with medications for migraines, I could figure out which drug is the best choice without as much trial and error.
Contrast this approach to what we do now with most genetic testing. In the current situation, a patient comes in with a specific diagnostic picture. For example, maybe the patient has Crohn’s disease but the pathologist is not sure based on the biopsy, or maybe there is a disorder of iron metabolism where we want to understand the long-term risks. Here, we are testing the blood for the presence of a single abnormal gene based on clinical suspicion. This is a much more straightforward approach than testing for and identifying 32,000 genes which may or may not have anything to do with the price of tea in China.
The average person probably
has 1,200 such single letter changes which cause
proteins to be slightly different.
And this brings us to the first big limitation in genomic medicine, the signal-to-noise ratio. We know that genetic defects can lead to catastrophic outcomes for patients, but the vast majority of genetic “defects” are harmless. The most common type of genetic change or defect is the single nucleotide polymorphism, or SNP (pronounced “snip”). This is when exactly one letter, A, C, T, or G, is substituted for another. If you switch an Adenine to a Thymine on chromosome 11 where hemoglobin is coded, then the resulting amino acid is going to come out as valine instead of glutamic acid. It is like a typo. If the recipe calls for salt but it mutates to malt, well that is going to change things. The result in this case is a hemoglobin molecule of a different shape. If both your mom and dad have this same genetic defect, you will be born with sickle cell anemia.
The average person probably has 1,200 such single letter changes which cause proteins to be slightly different. Craig Venter, PhD, the biologist who first sequenced the entire human genome, found 1,200 mutations when he sequenced his own genes. Many of these changes are irrelevant to the functioning of a protein, but under the right circumstances, a few could be highly significant, yet we have no way to know.
The next big limitation in genomic medicine is that genes are only part of the story. A variety of diseases require not only a genetic predisposition, but also an environmental trigger. For example, in type I (juvenile onset) diabetes, if one sibling in a pair of identical twins is diabetic, the other will eventually develop type I diabetes about 80% of the time if you follow the twins for many decades, but not 100%. In schizophrenia, the rate is around 50%. So knowing someone’s exact genome is not enough to predict what diseases they will get. With conditions like asthma and heart attacks, the influence of environment is likely much greater.
Finally, only a small minority of the common diseases which affect our population are caused by single genetic defects. High cholesterol, depression, breast cancer, and migraines are likely influenced by many different genes. Given 32,000 different genes and 1,200 “defects” in a normal healthy person, the task of associating a genetic defect with a particular disease is tough. Considering that most diseases are caused by the interaction of dozens of different genes with the environment, we can see that even having a patient’s entire genome sequenced and compared to a database of thousands of other people is going to make it very difficult to extract useful information.
A perfect example of this problem comes from the company 23andMe which will do a lot of genetic testing on your saliva for $99. Several patients have had this analysis performed and sent me the results. Amazingly, some of my Caucasian patients whose parents came from Europe found out through sophisticated genetic testing that they are indeed white people. But most of the reporting came in as an increased or decreased risk of certain diseases. For example, 1.6% of the population overall develops Parkinson’s disease, but with a single letter change in the PARK8 gene, the risk rises to 2.2%. Well, if you’re selling genetic testing, you can say someone’s risk went up by 38%, but the reality for most of us is that the difference between a 1.6% and a 2.2% risk of Parkinson’s is not meaningful. In addition, many patients have quite mild Parkinson’s and only a few have very severe disease. This problem underscores how little we know about the ways that genetic changes cause most of the common diseases that people suffer from.
Genomic medicine has great promise, but the devil is in those billions of little details. I predict that success is going to come slowly and incrementally. At this point, it is a great research tool that is not quite ready for prime time in clinical medicine, but its day will come eventually.

